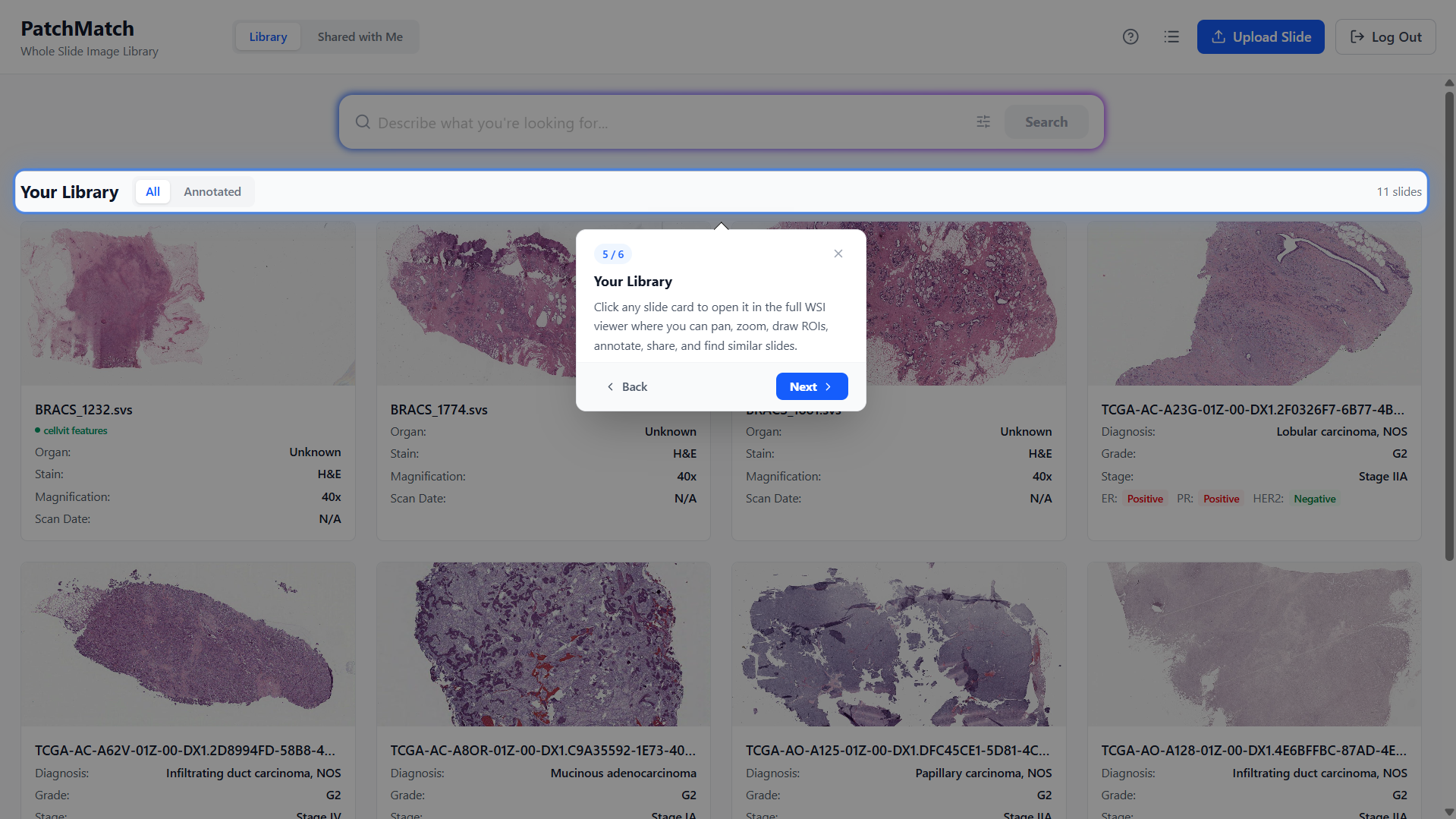
Screenshots from
the PatchMatch workflow
A visual walkthrough of the application.
Login Page
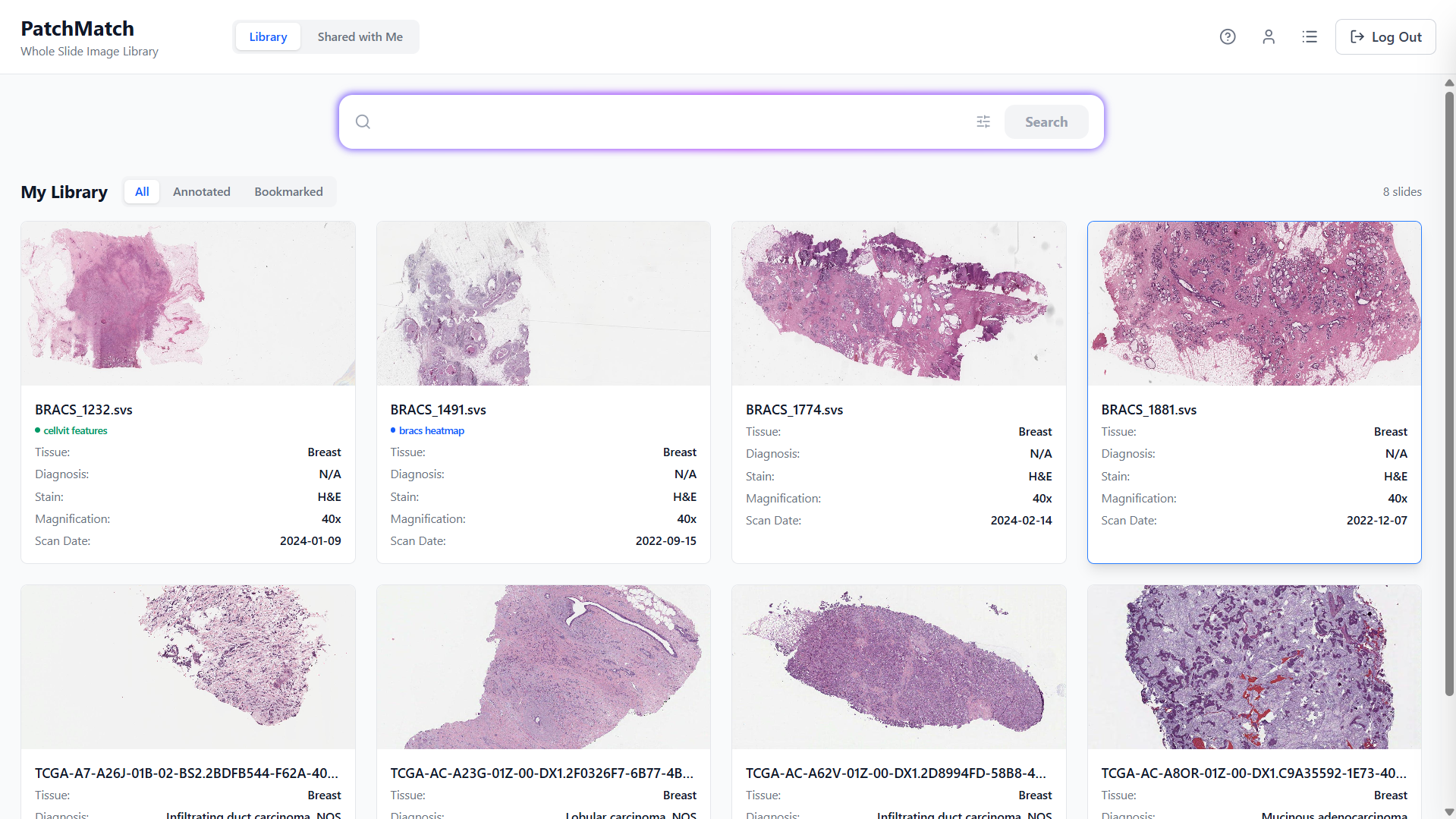
Library Page
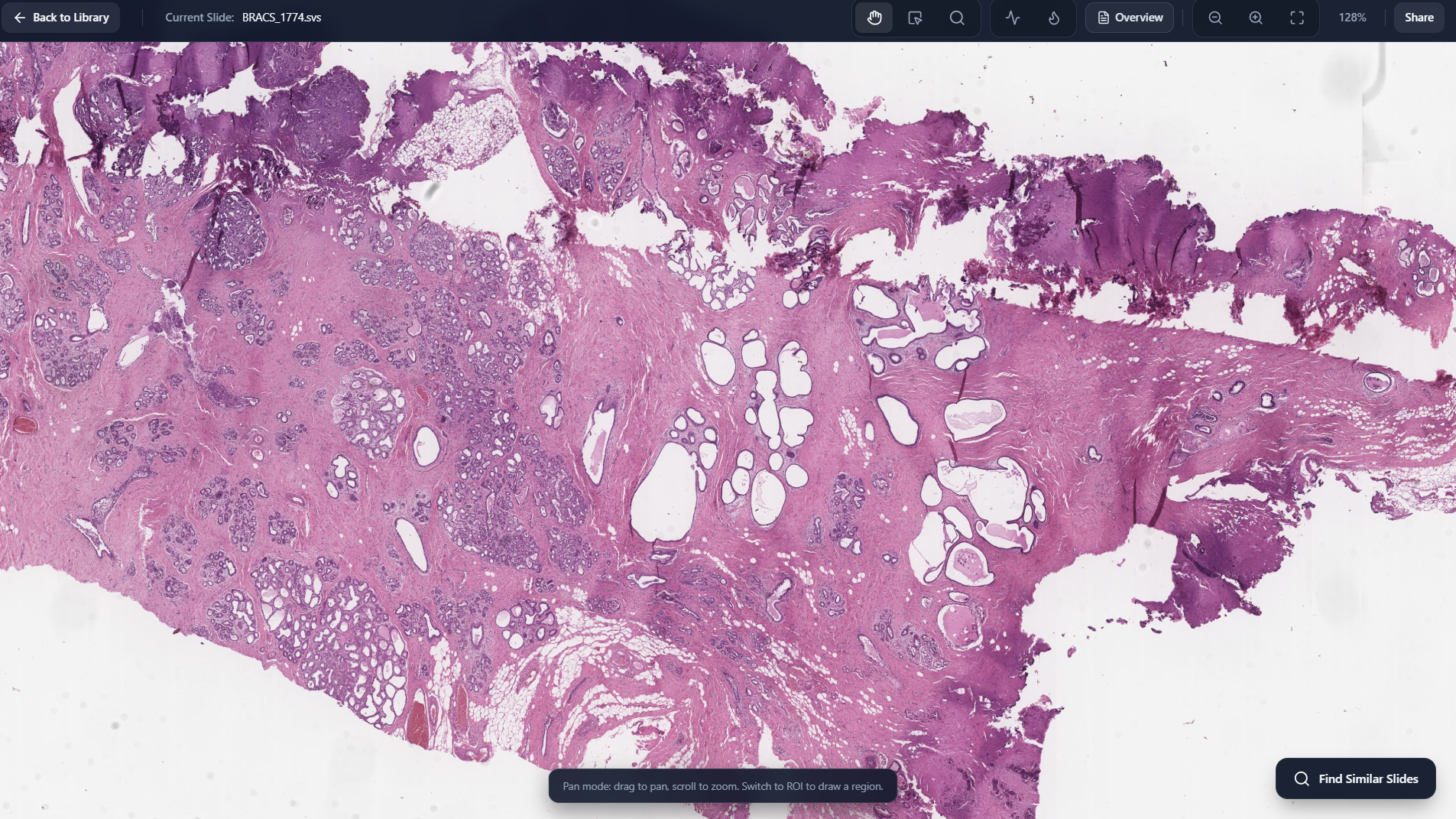
WSI Viewer
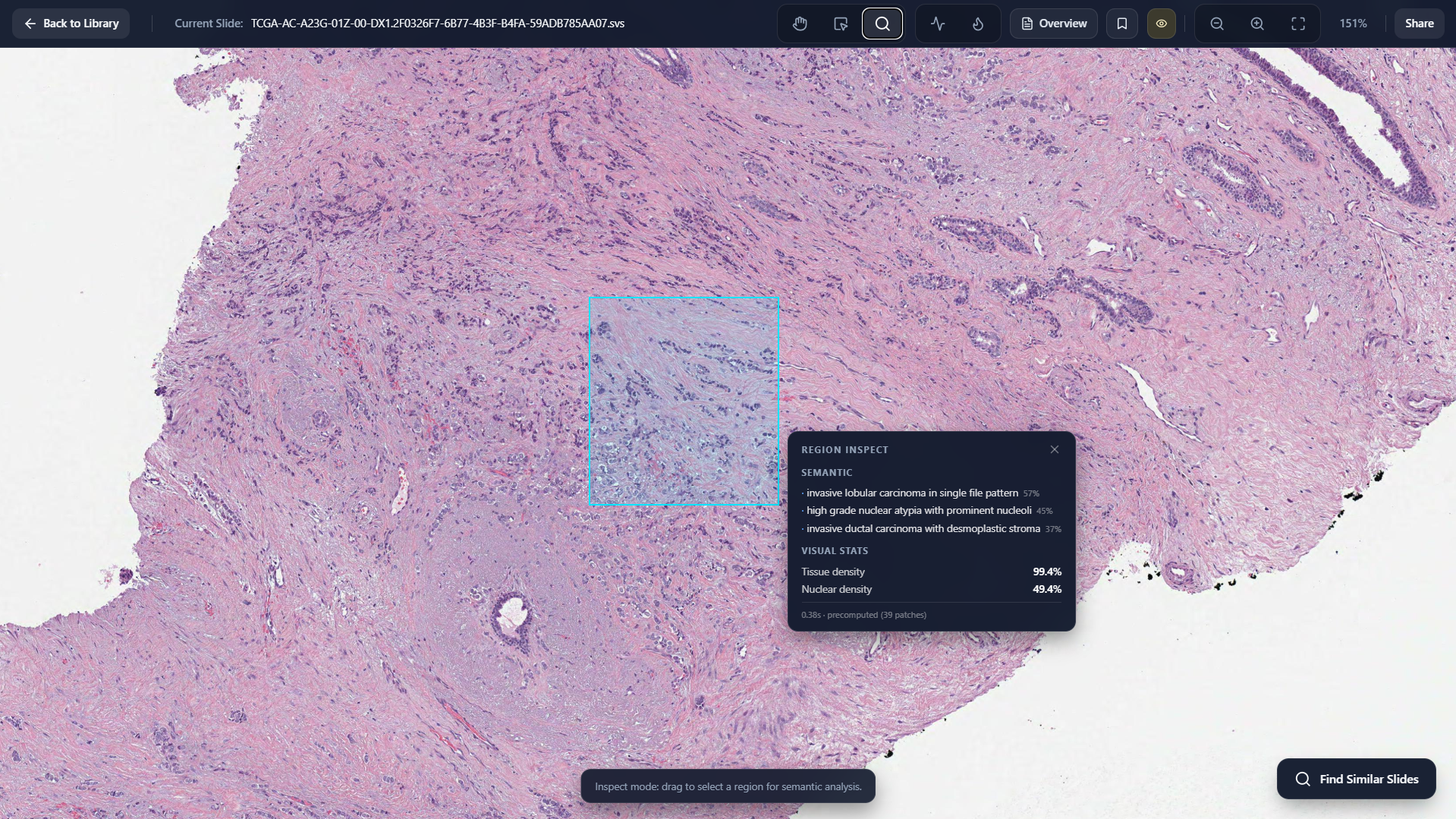
Inspect Region
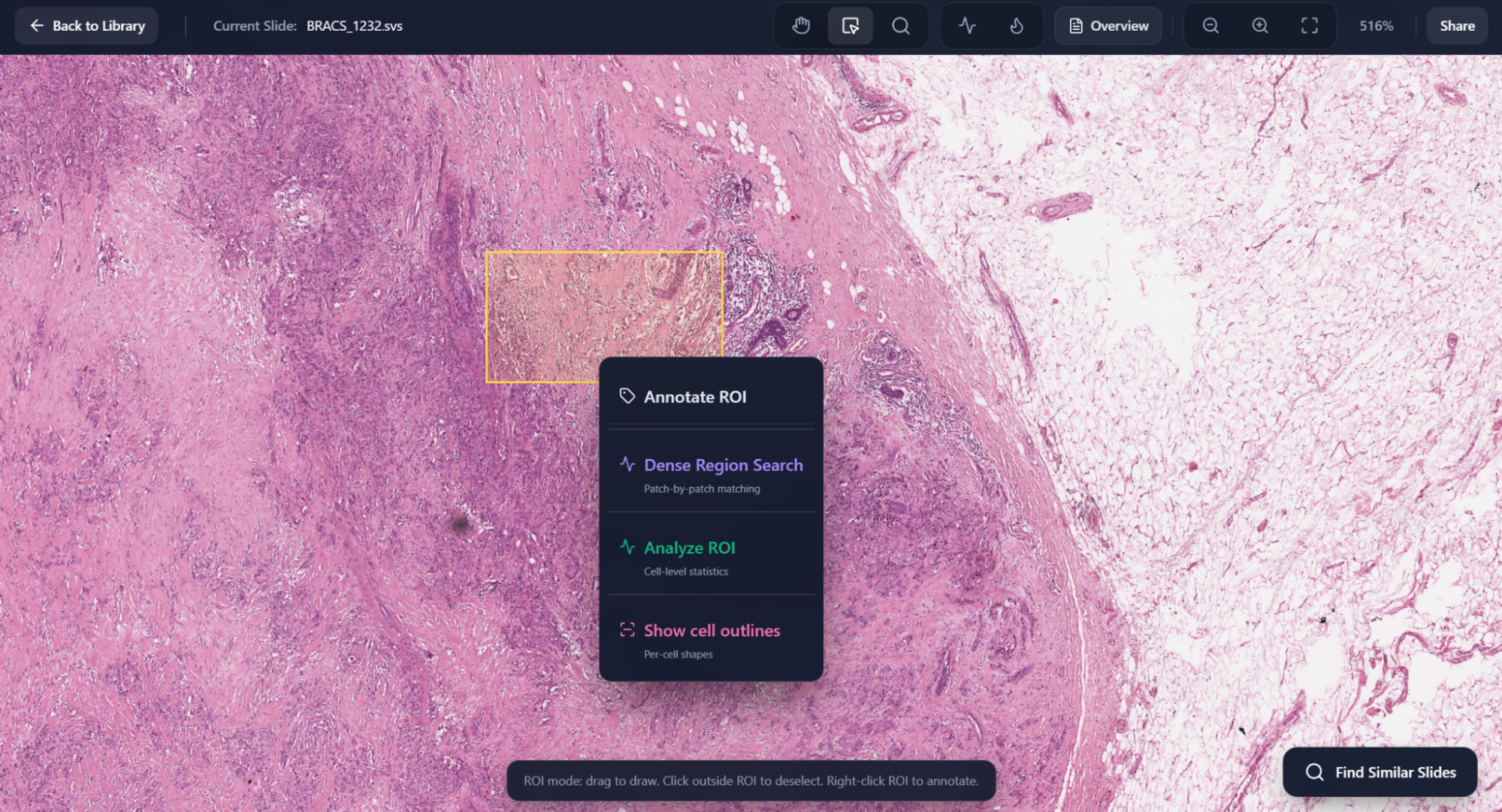
Select Region
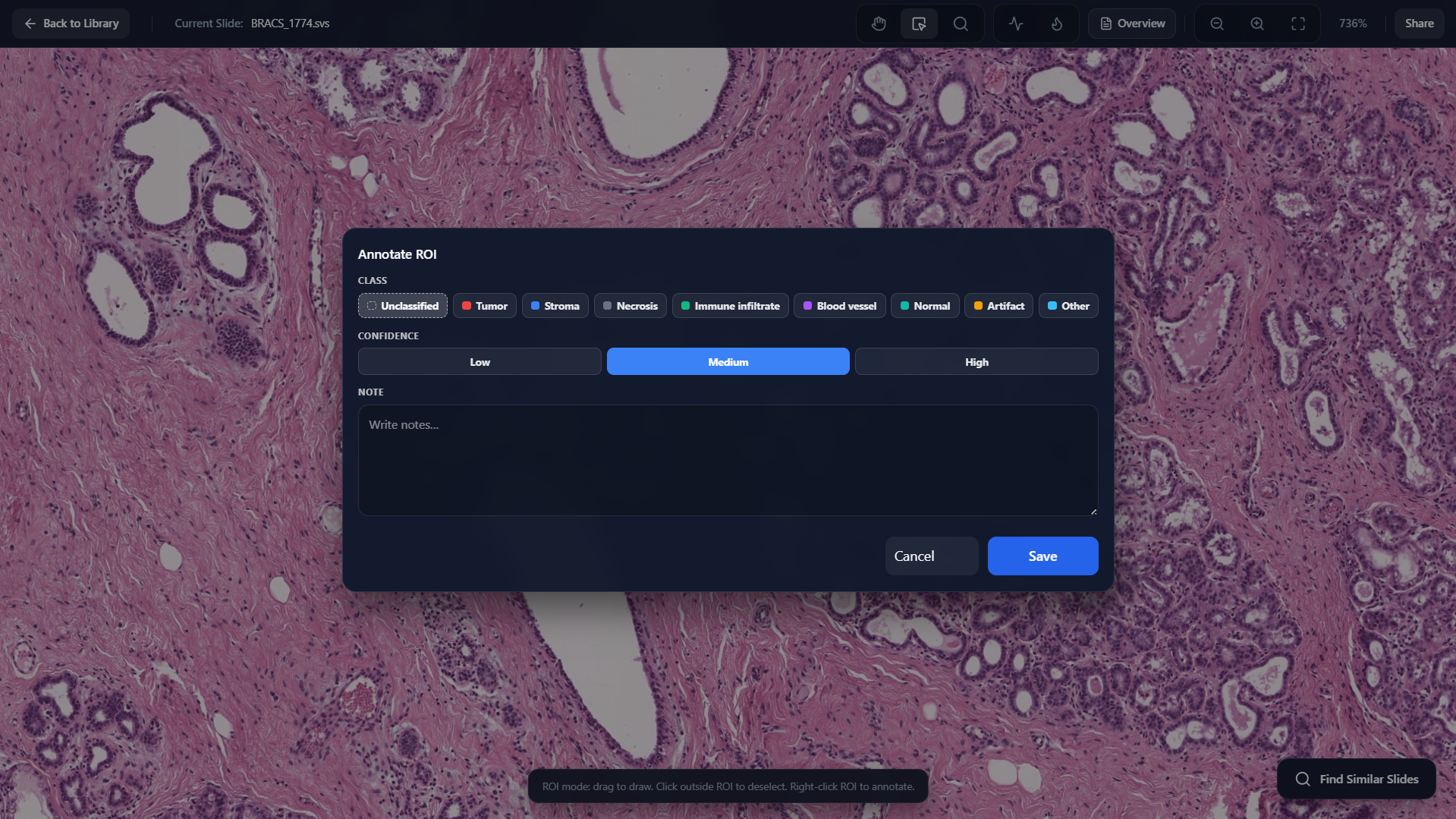
Annotate Region
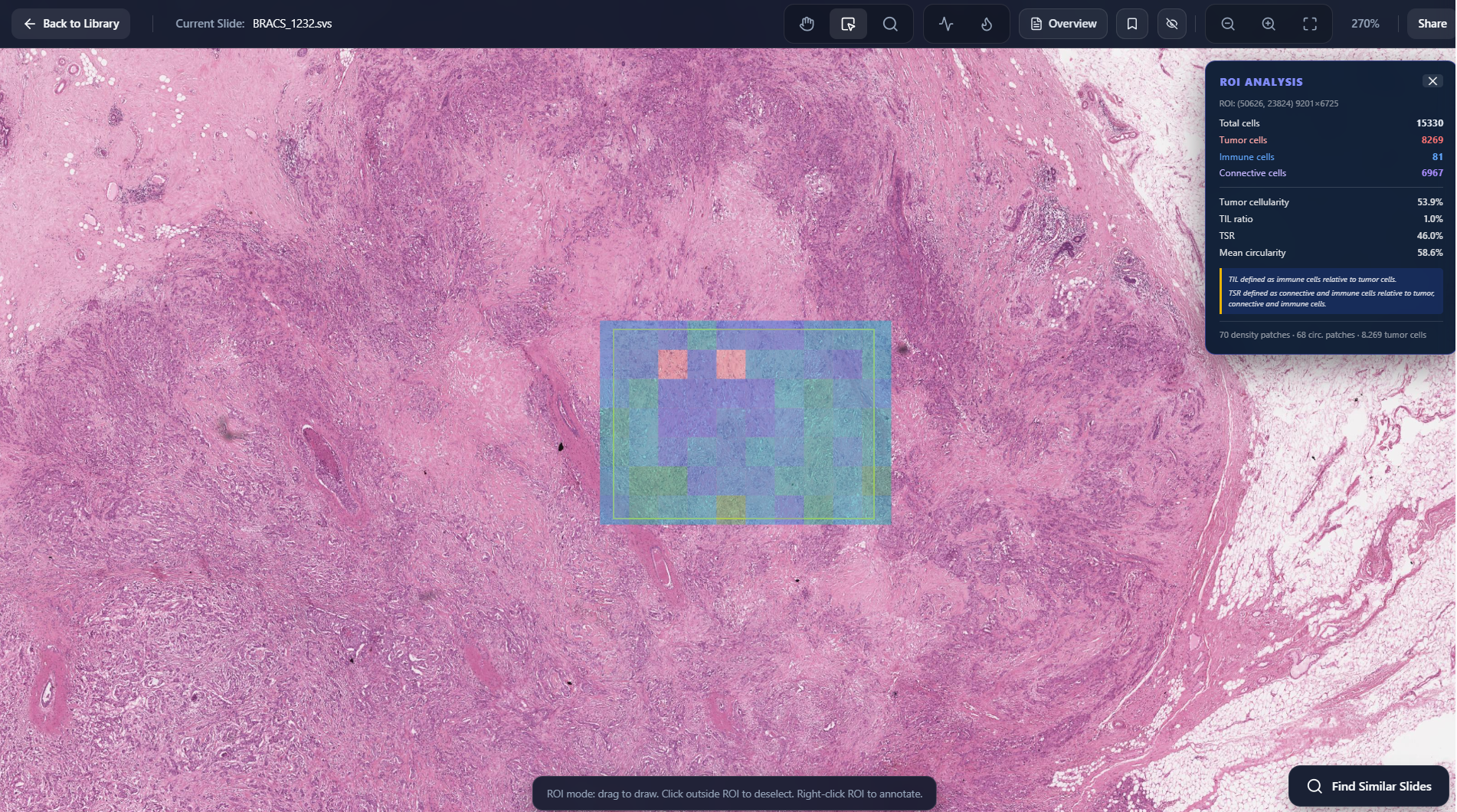
Region Level Cellular Metrics
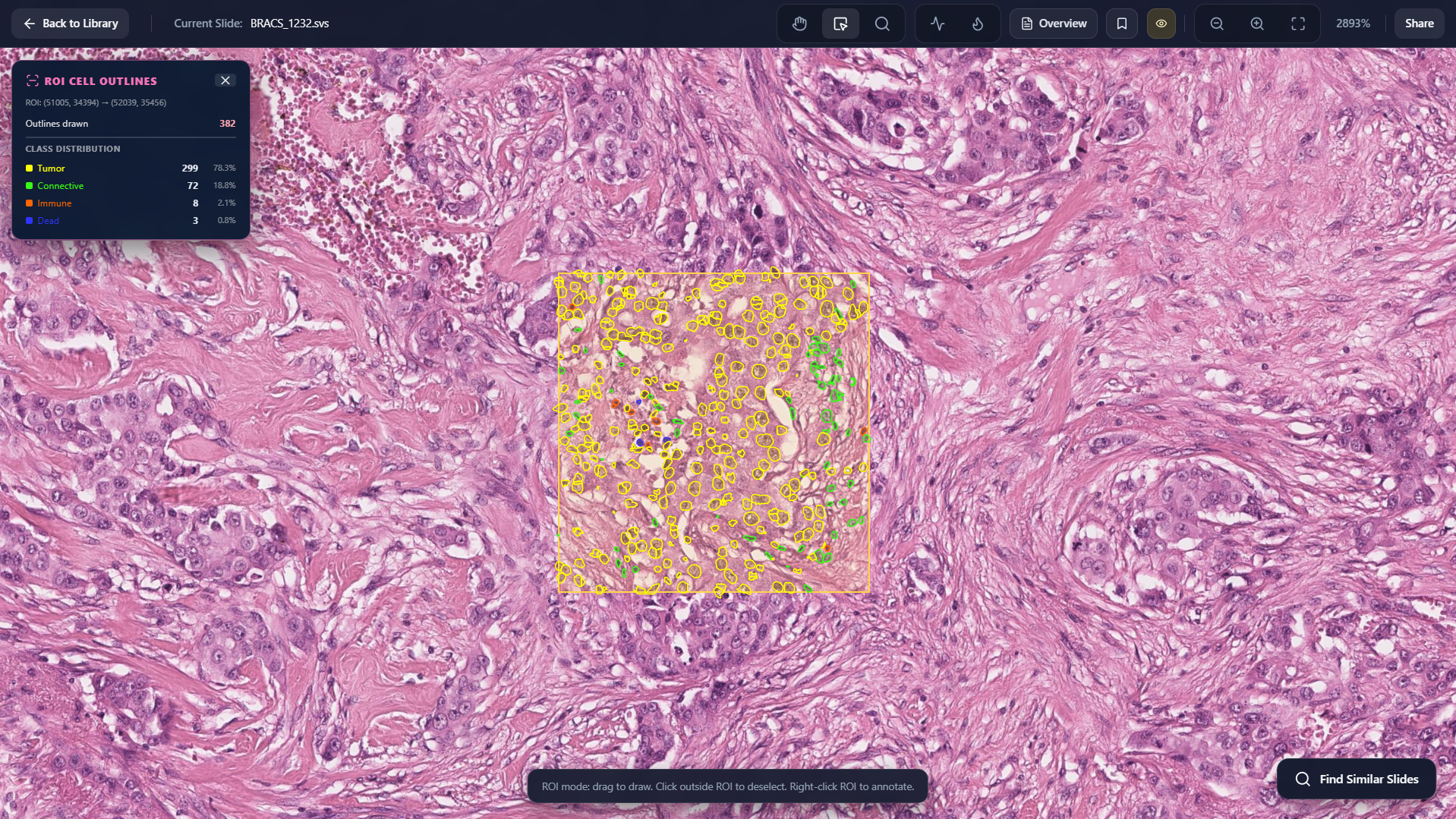
Cell Outlining
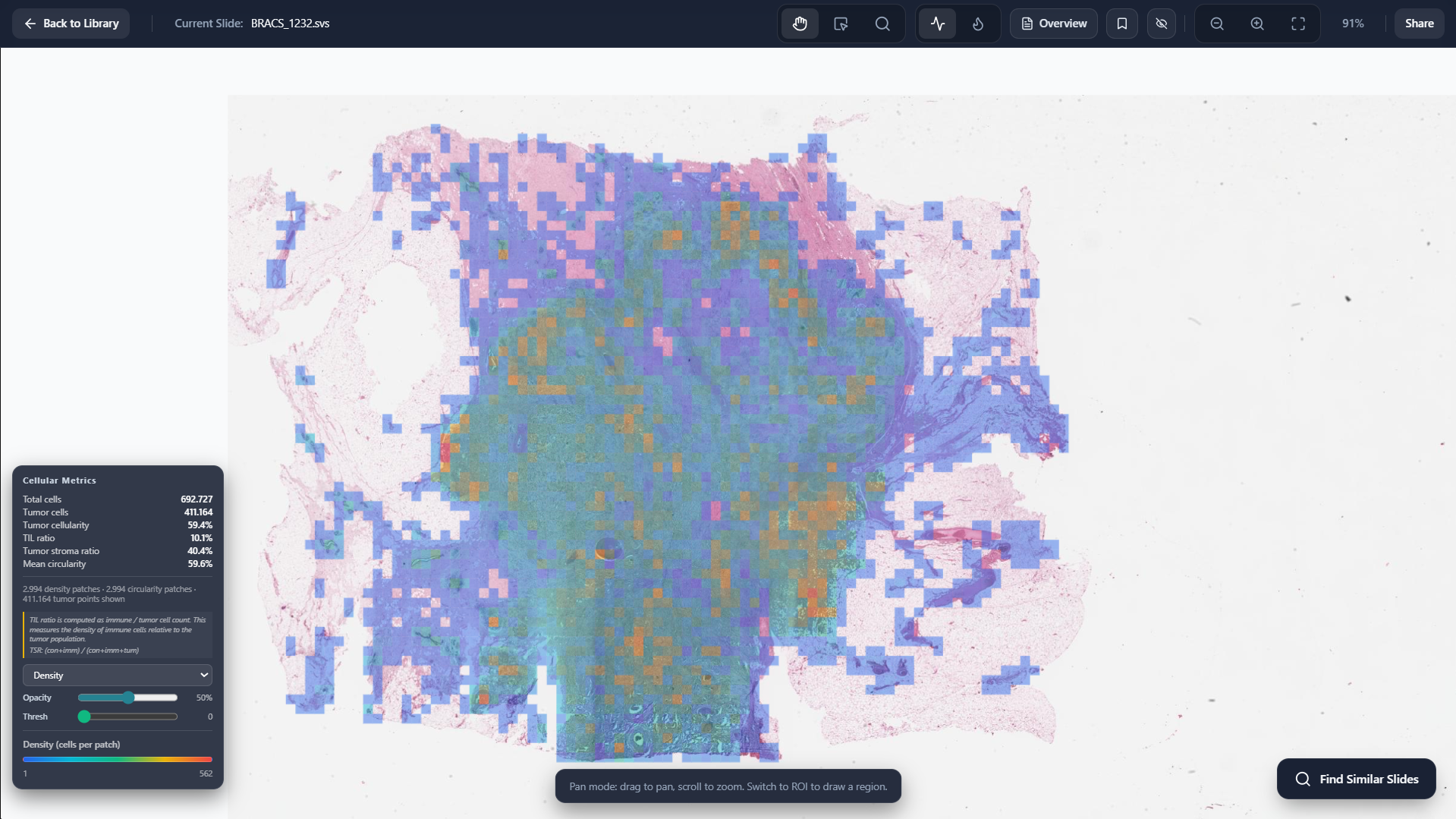
Slide Level Cellular Metrics
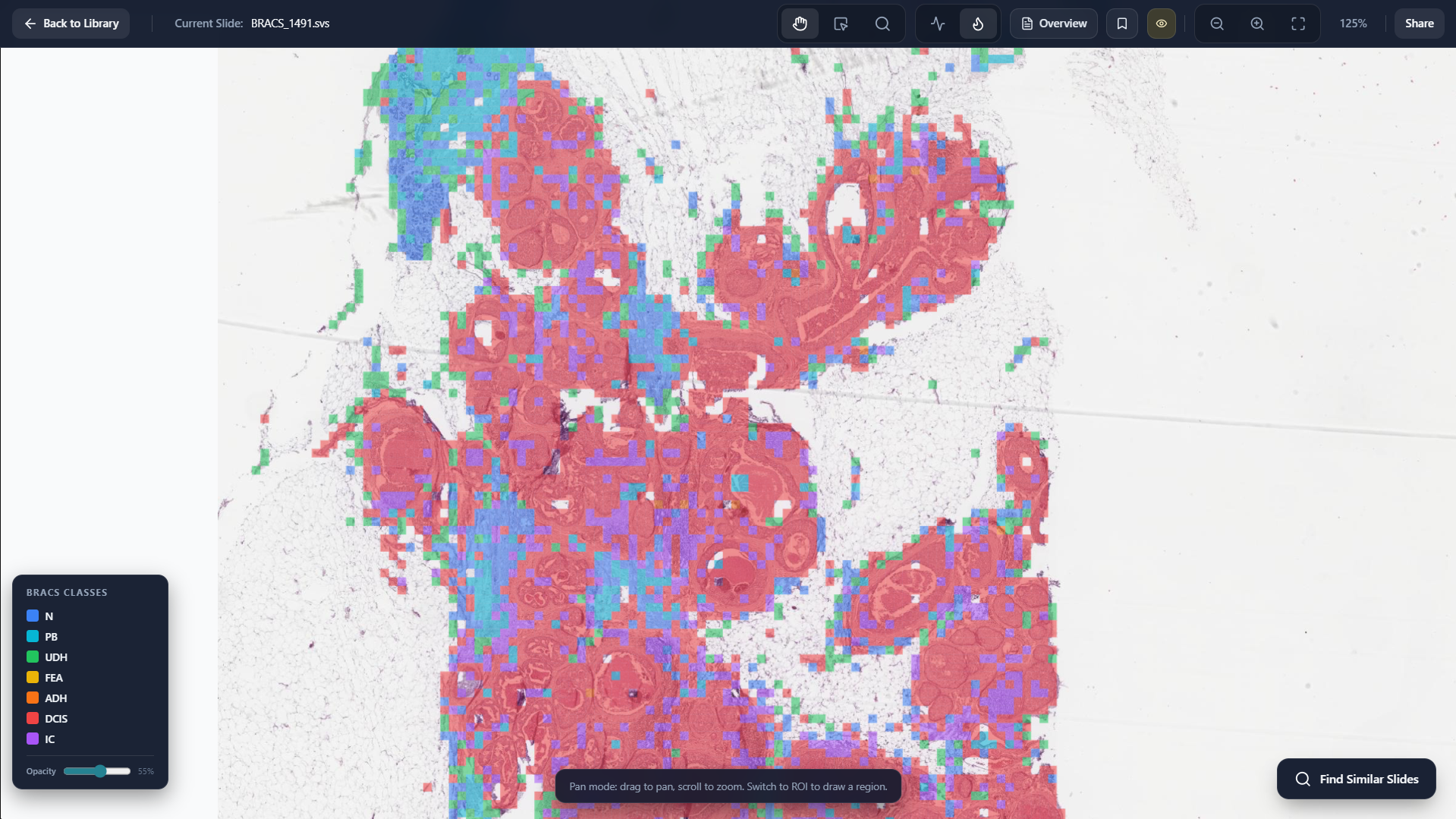
Class Heatmap
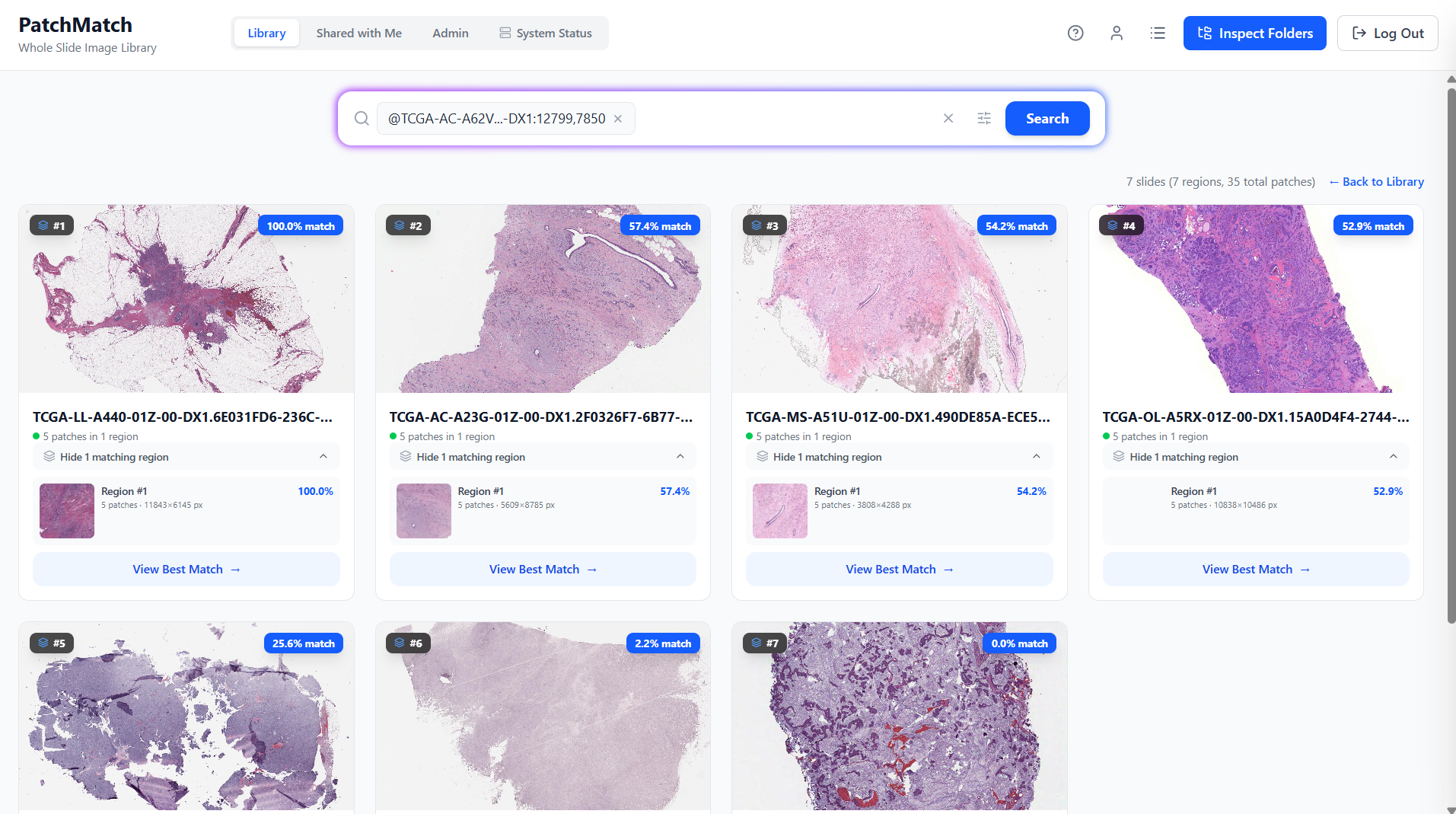
Region To Slide Retrieval
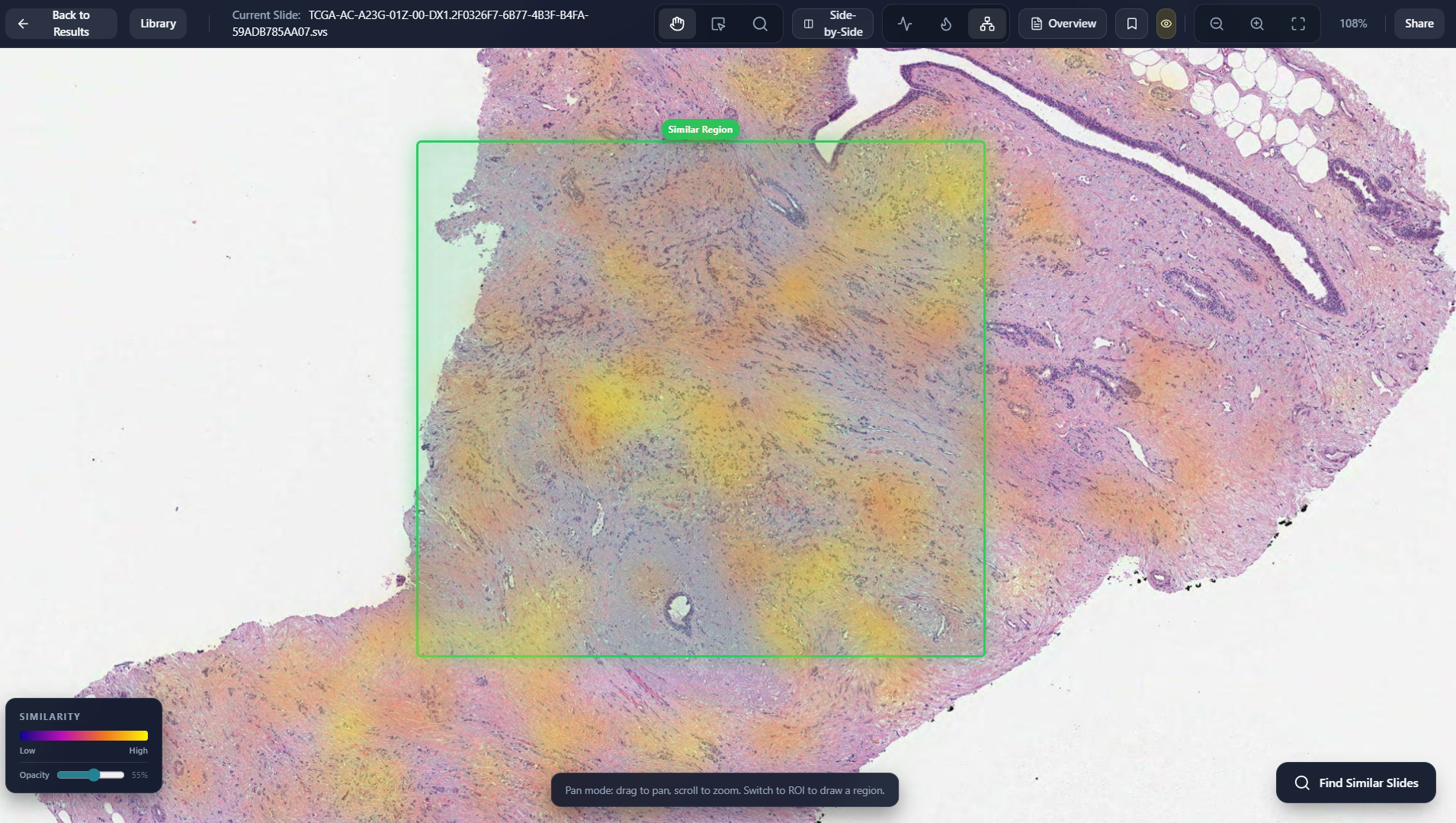
Display Retrieved Region
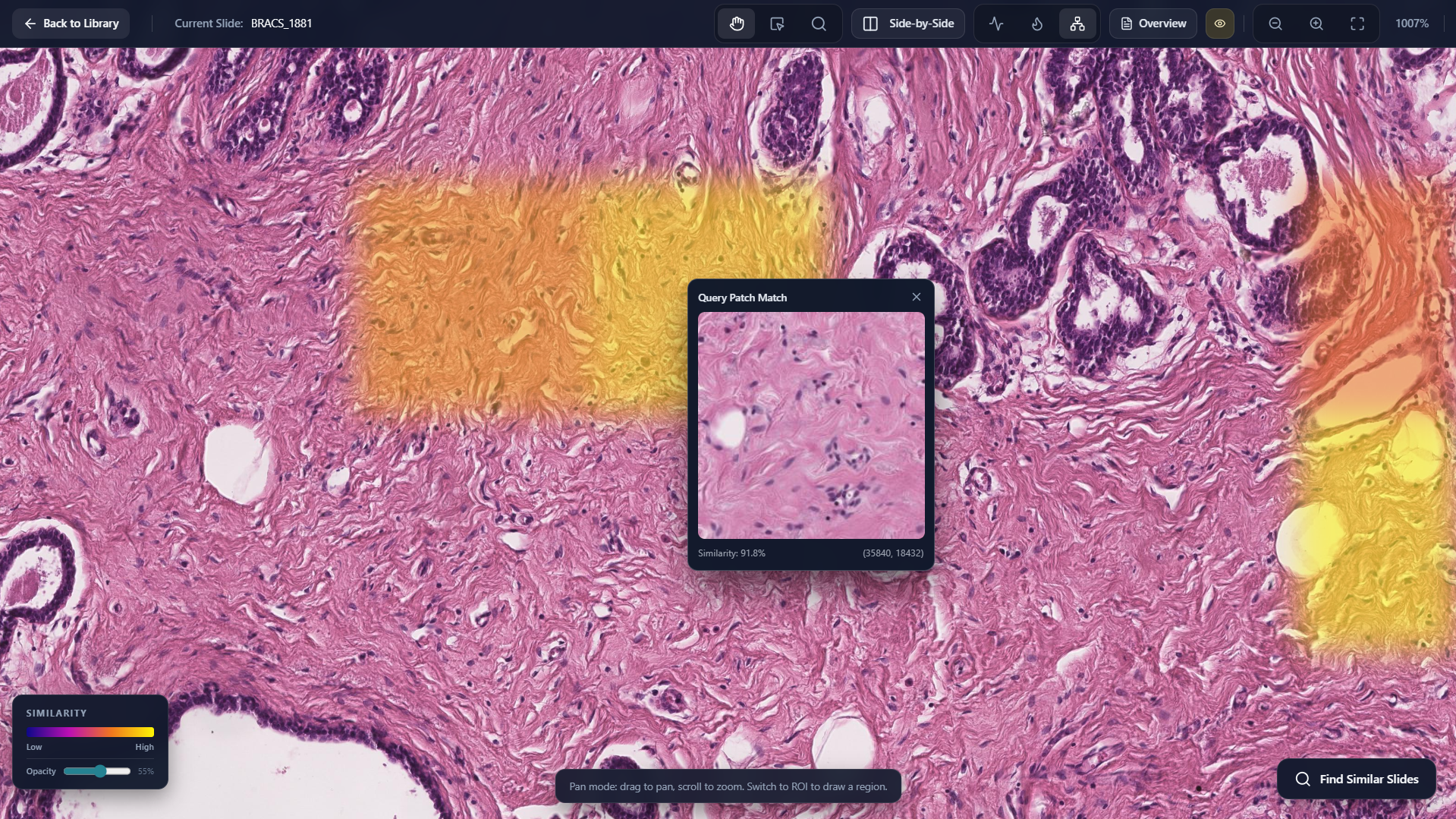
Display Matching Query Patch
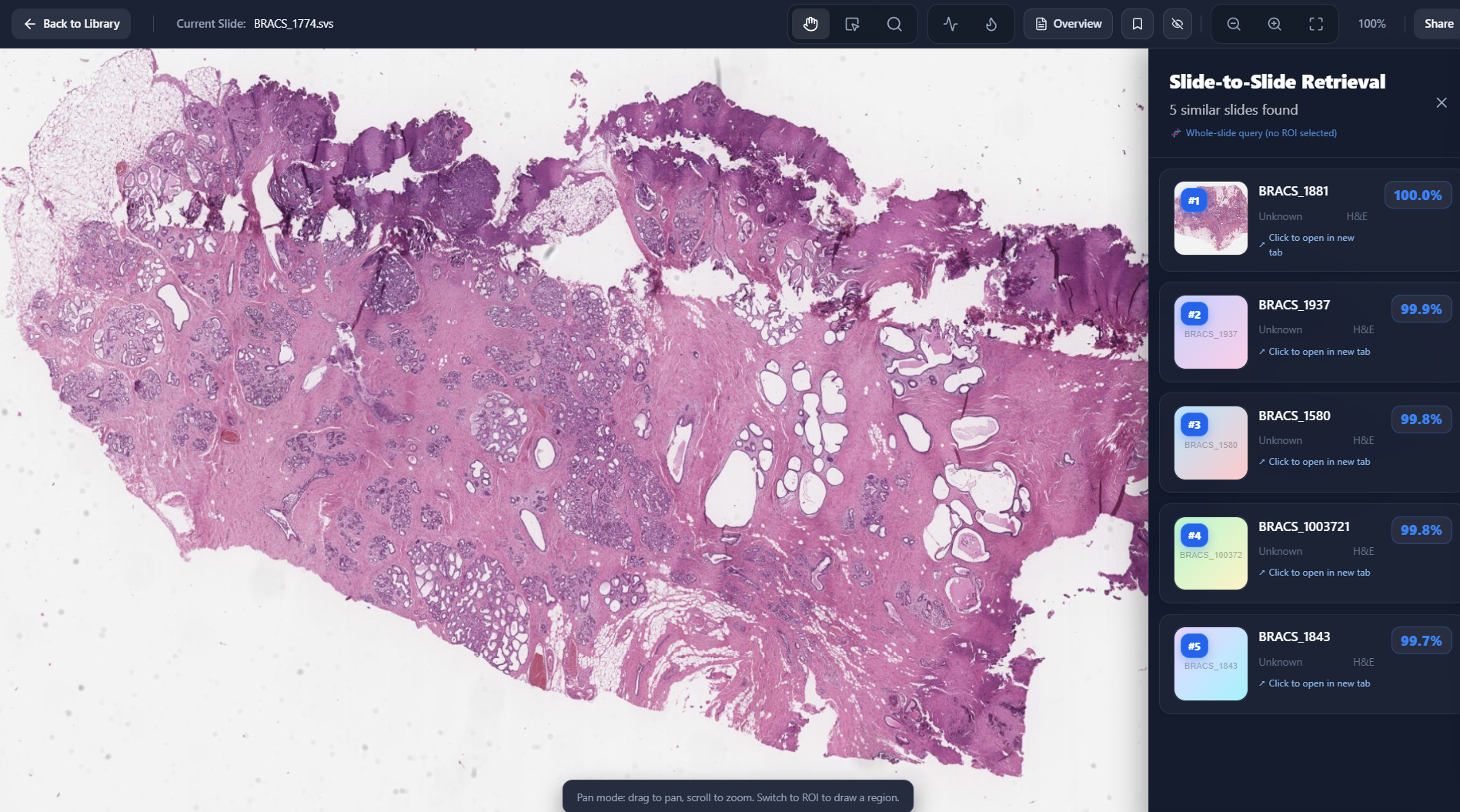
Slide To Slide Retrieval
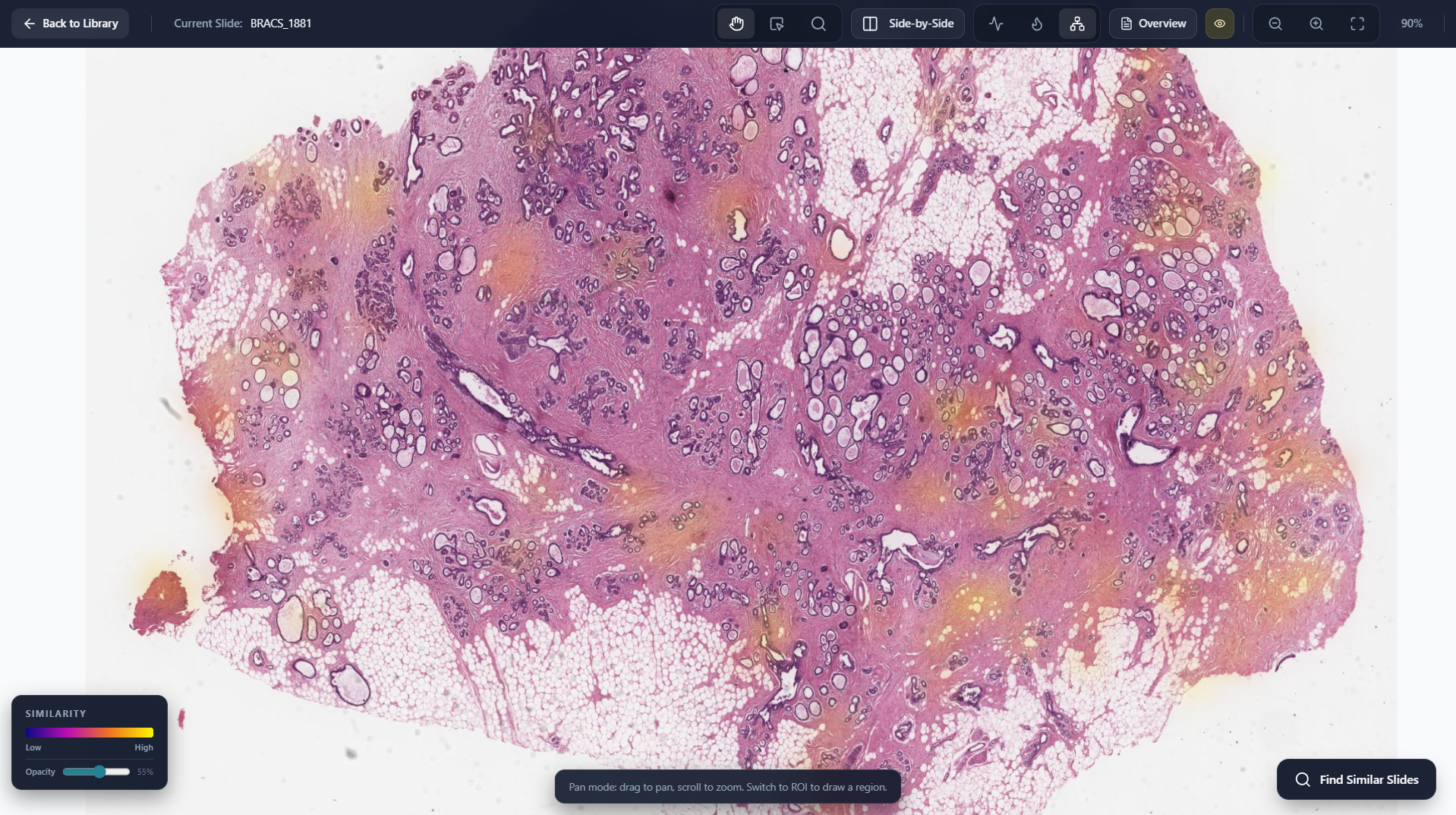
Similarity Map
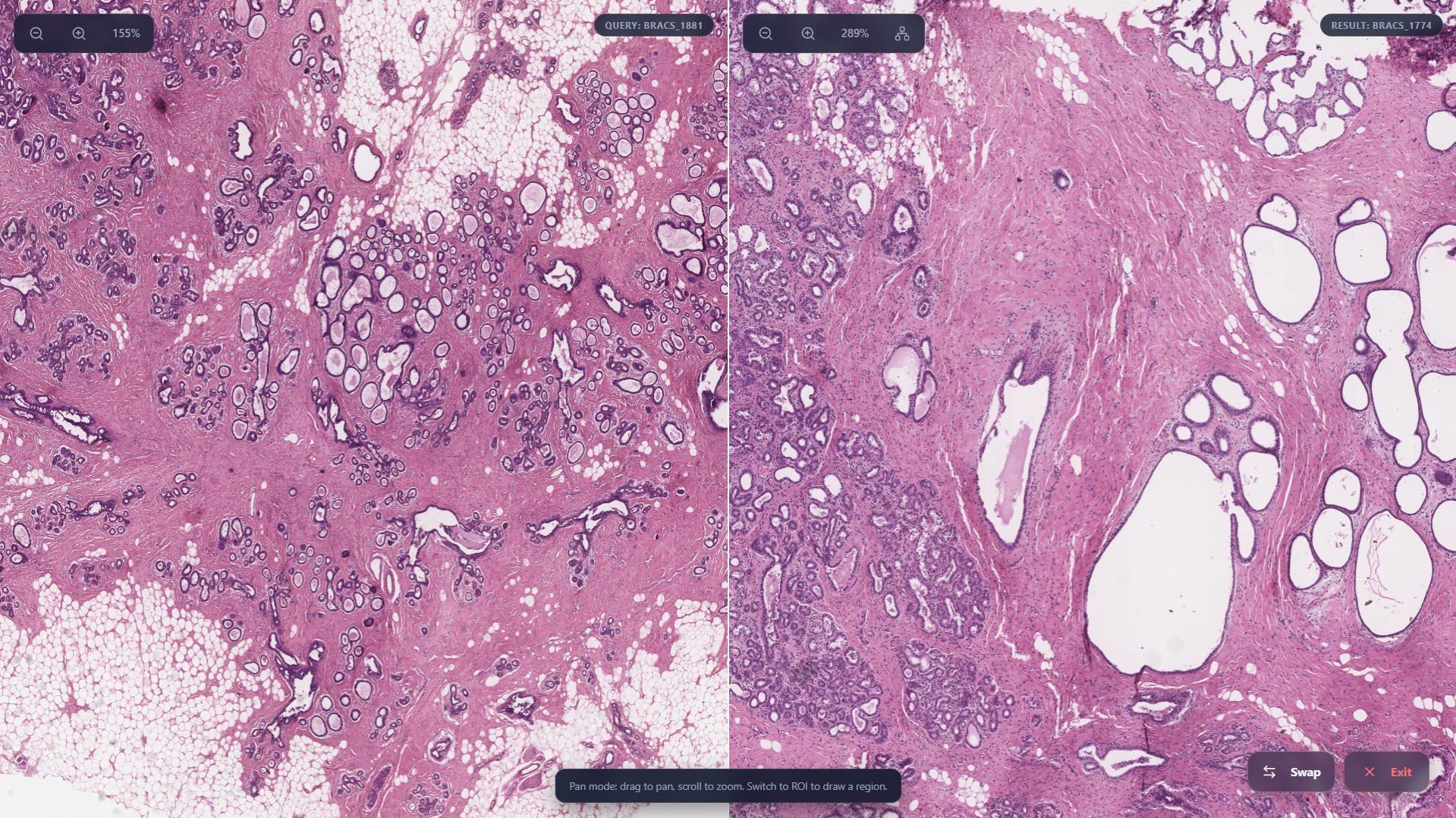
Side By Side View
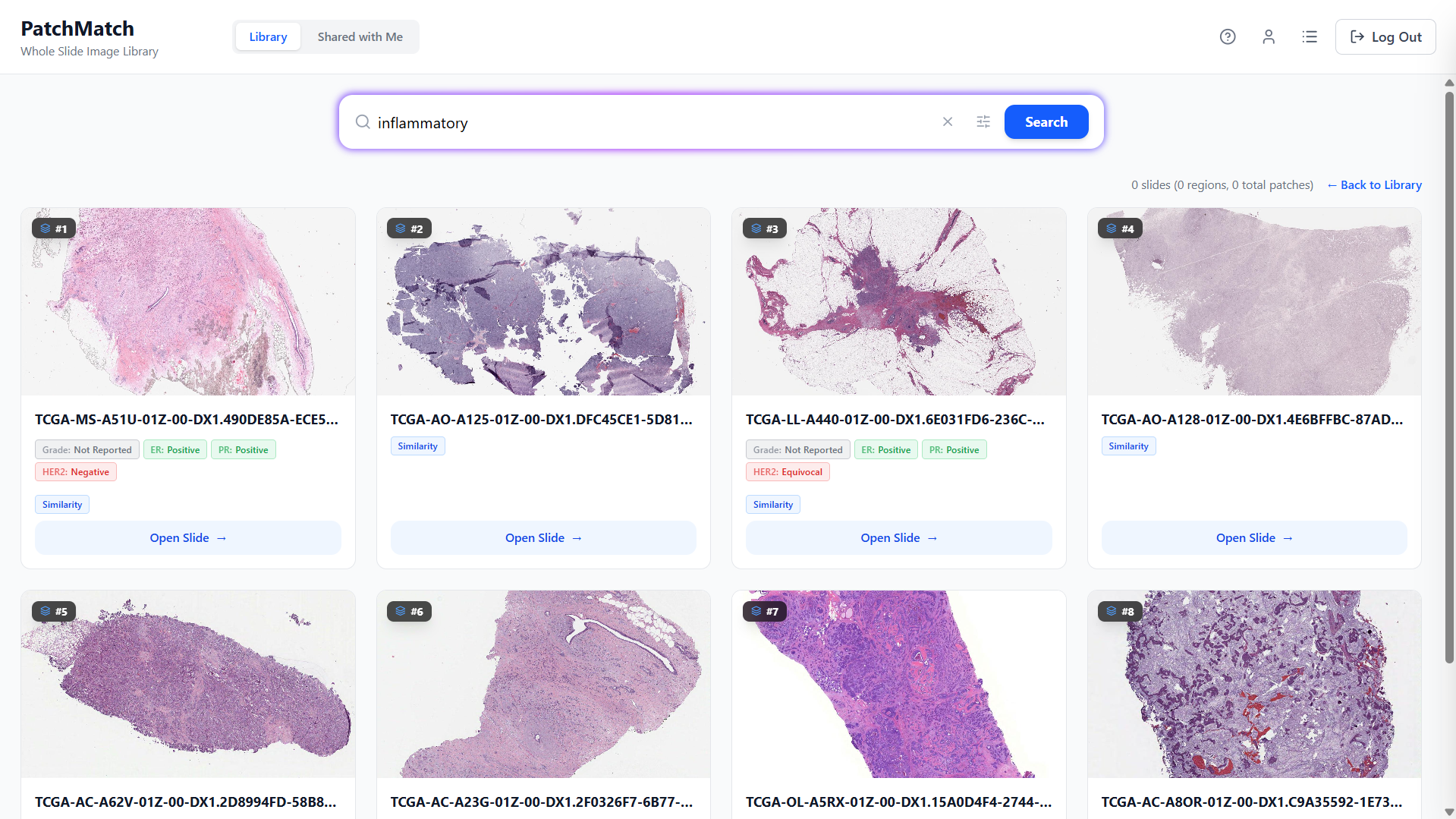
Text To Slide Search
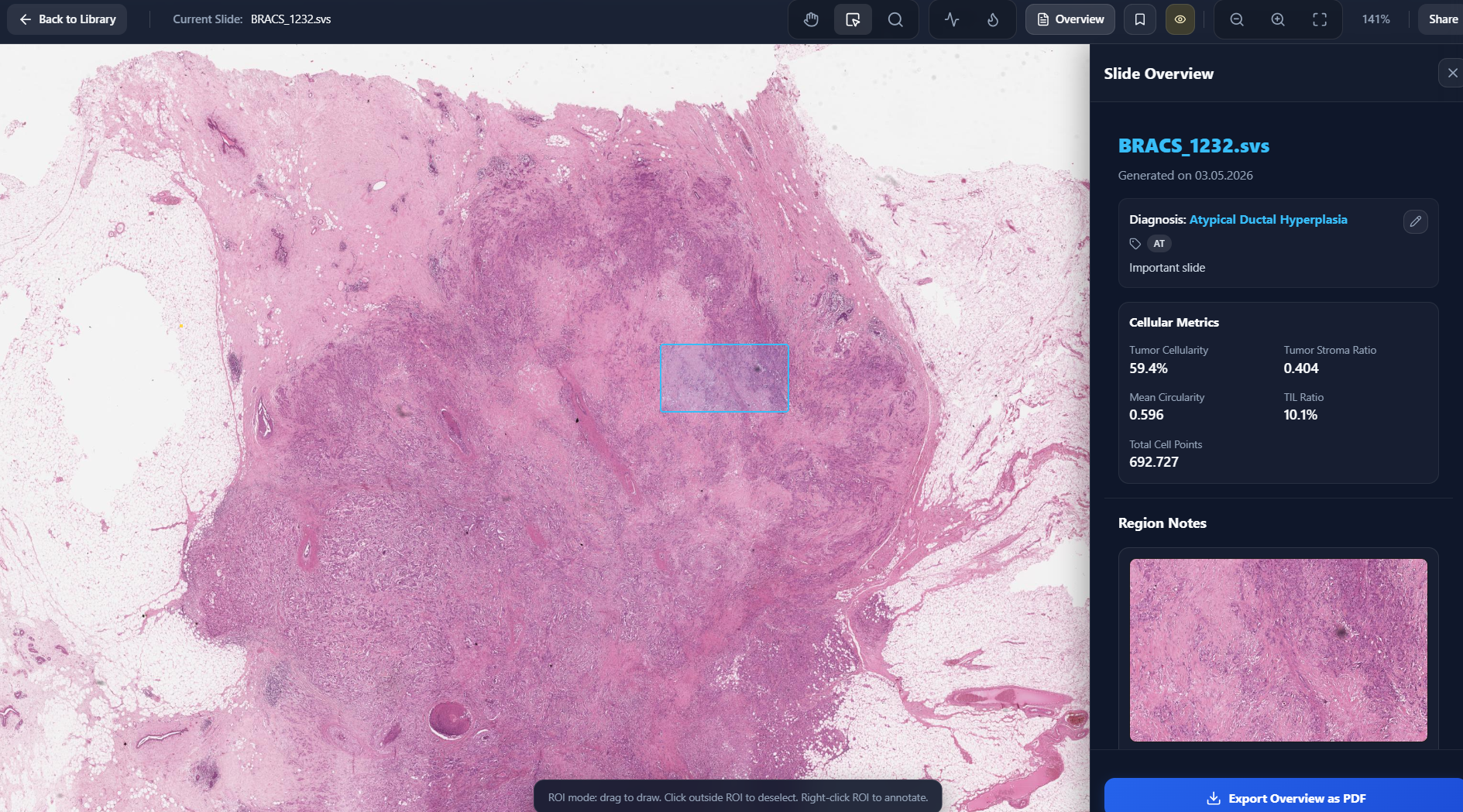
Slide Overview
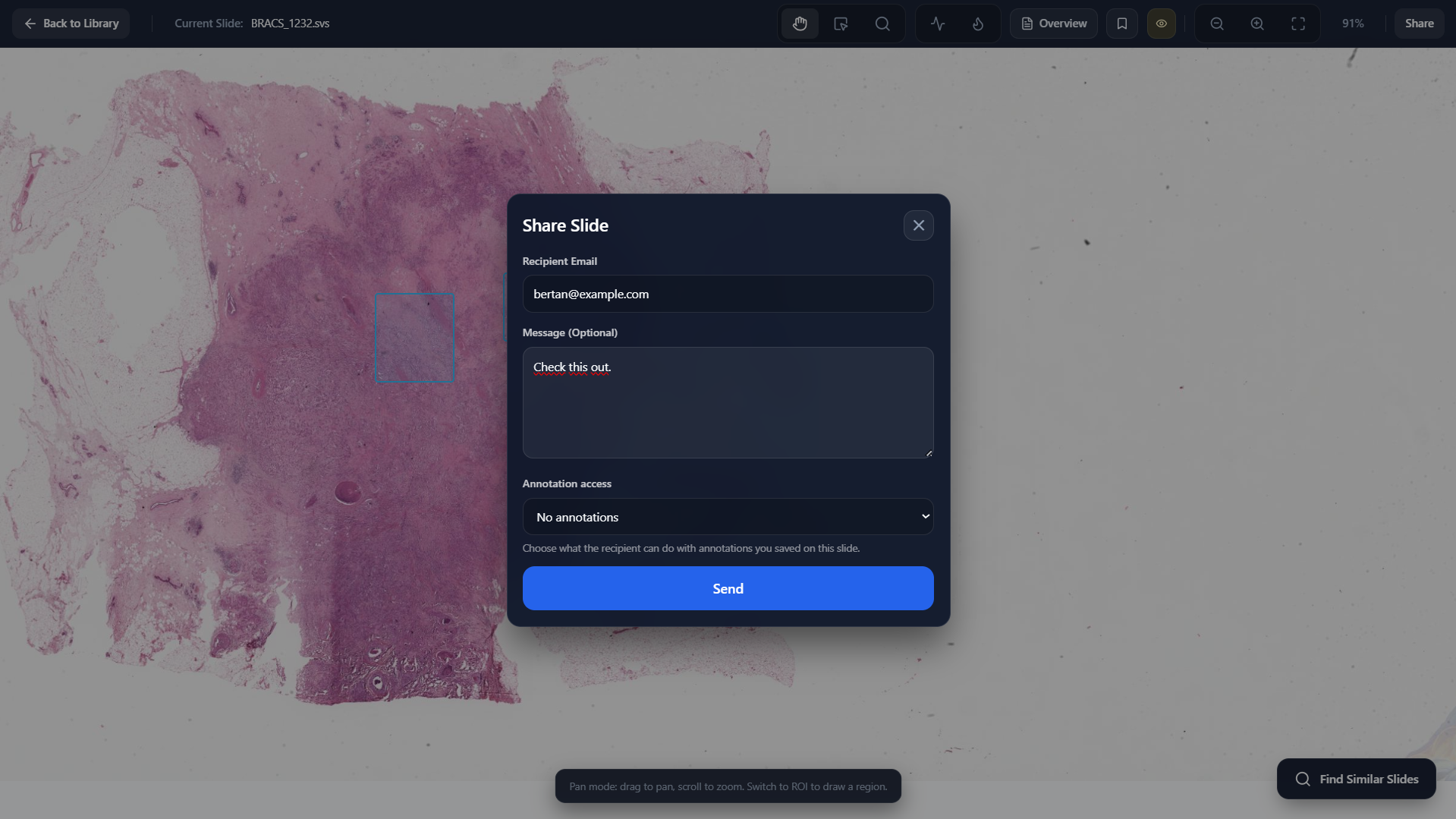
Share Slide
Shared With Me Page
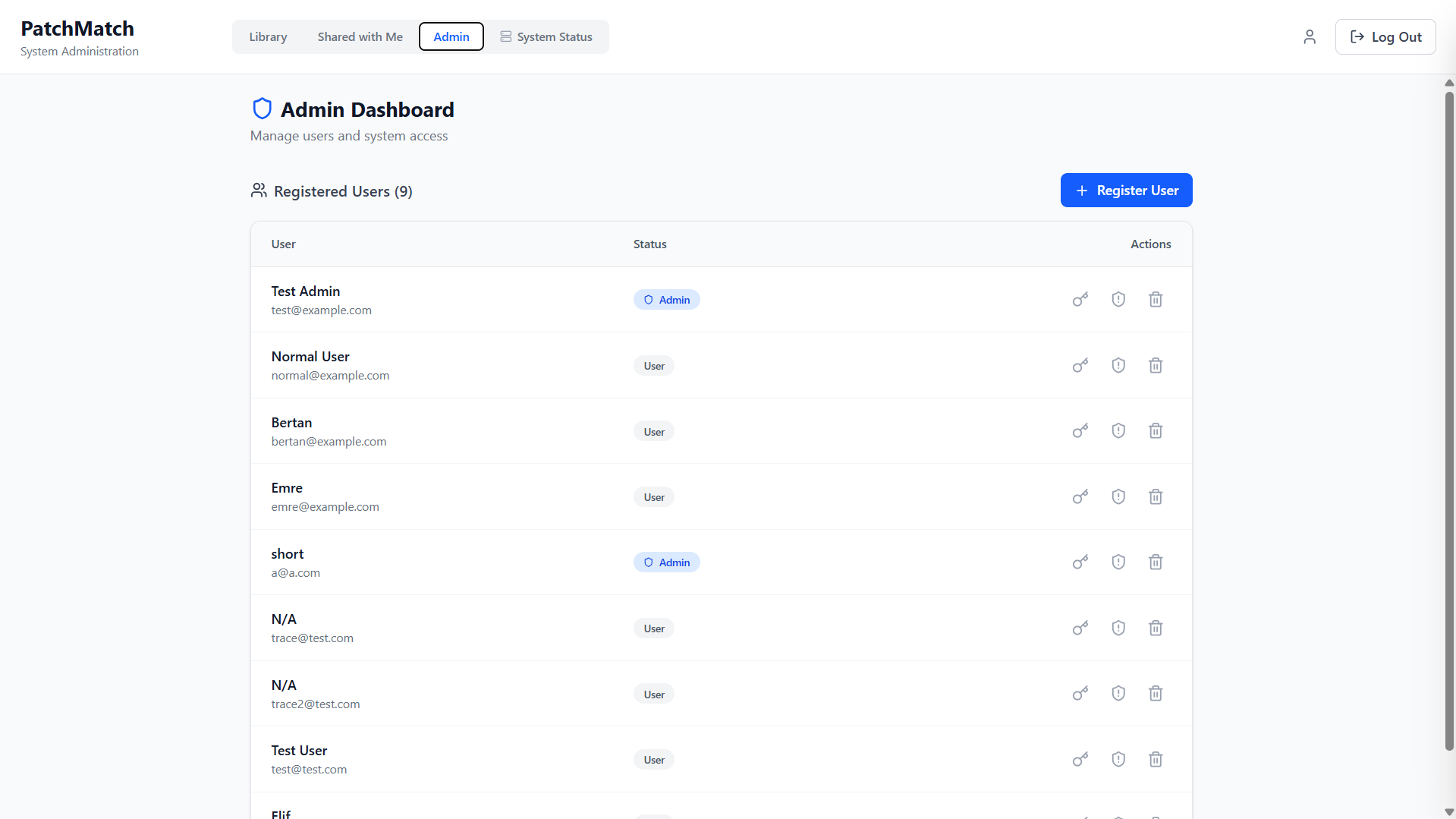
Admin Dashboard
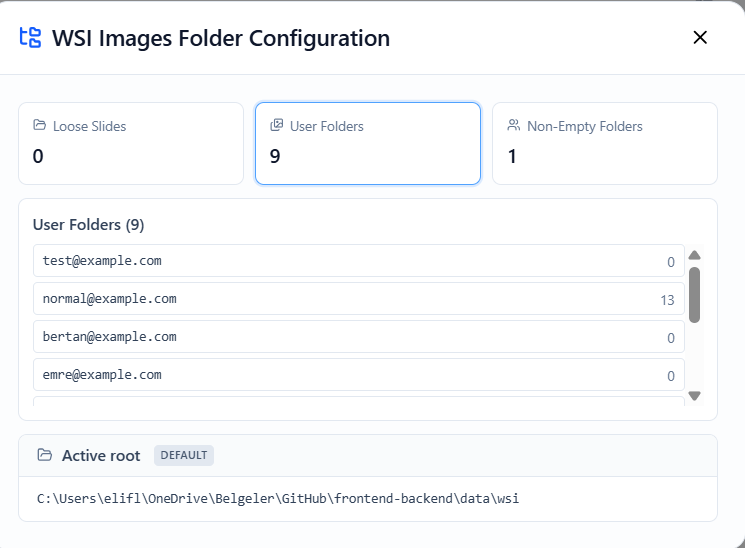
Inspect Folders
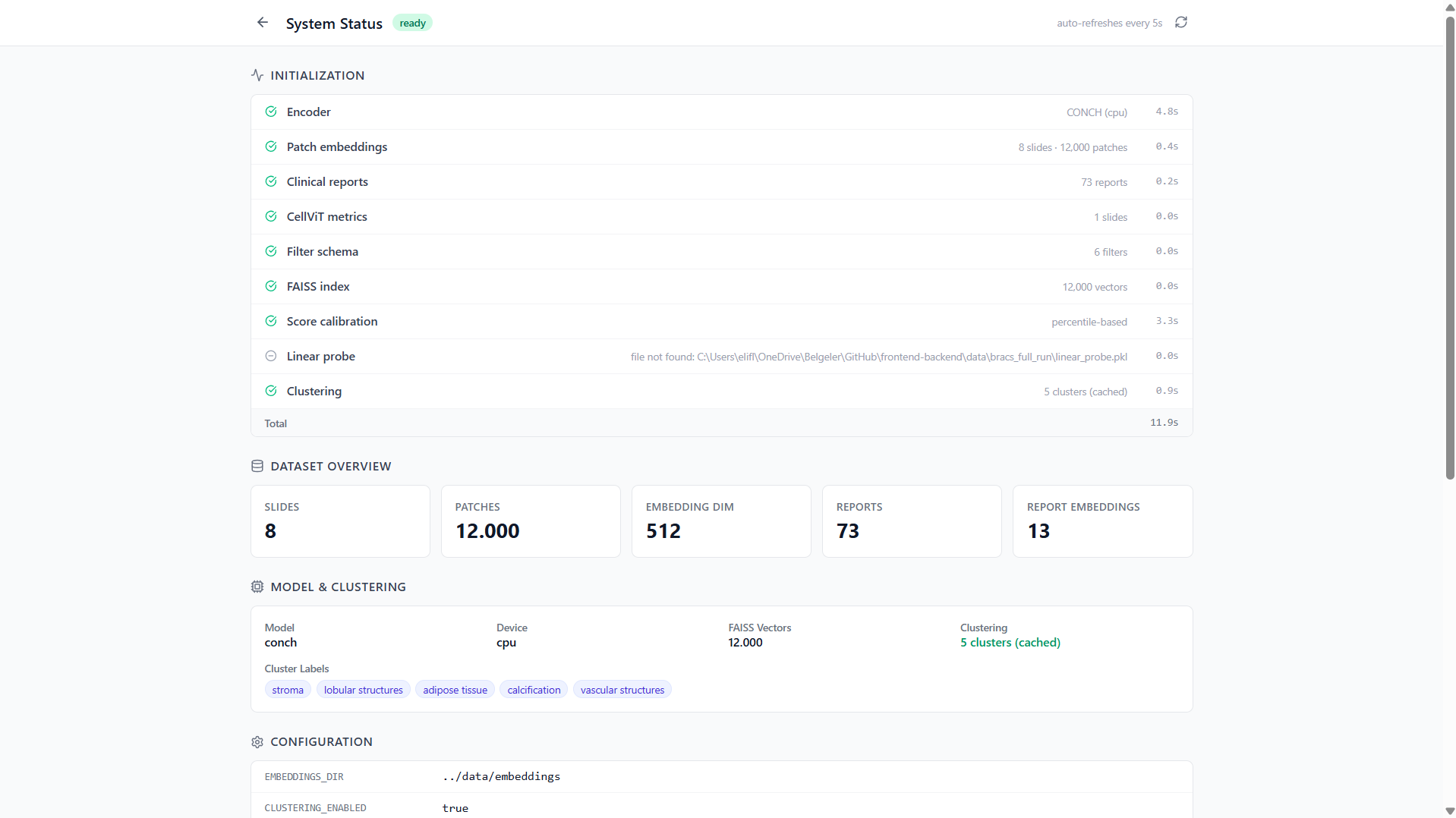
System Status Page